Overview
Microarray dataset is a series of microarray experiments, in which each experiment represents a gene expression signature of a tissue sample. Typical dataset has a fairly small sample size, usually less than one hundred, whereas the number of genes involved is more than several thousands. Cluster analysis of microarray gene expression data is aimed at finding subclasses of disease at the molecular level. In view of statistics, one major difficulty in this problem is that the number of samples to be clustered is much smaller than the dimension of data which is equal to the number of genes involved in an experiment. Under such a situation, model-based clustering according to a conventional finite mixture model might fail due to the occurrence of overfitting during the density estimation process. The mixed factors analysis was originally developed to overcome such difficulty in microarray gene expression profilings.
The mixed factors model presents a parsimonious parameterization of Gaussian mixture model. Consequently, the method enables us to avoid the occurrence of overfitting even when the dimension of data is more than several thousands! The method contains the following applications:
- Clustering microarray experiments
- Data visualization via the built-in dimension reduction system
- Identification of module transcriptional genes that are relevant to the calibrated clusters
- Determination of an appropriate number of clusters
- Determination of the number of module transcriptionals
- Missing data imputation
For a belief introduction of the mixed factors analysis, see What's the Mixed Factors Analysis ?. For more details, see our papers, e.g. Yoshida et al. (2004).
- Home
- Overview
- Software Description
- What is Mixed Factors Analysis?
- Online Manual
- ArrayCluster Developers Info.
- Citation (under construction)
- Bug Tracking (under construction)
Figure 1: Snapshot of graphical user interface of ArrayCluster
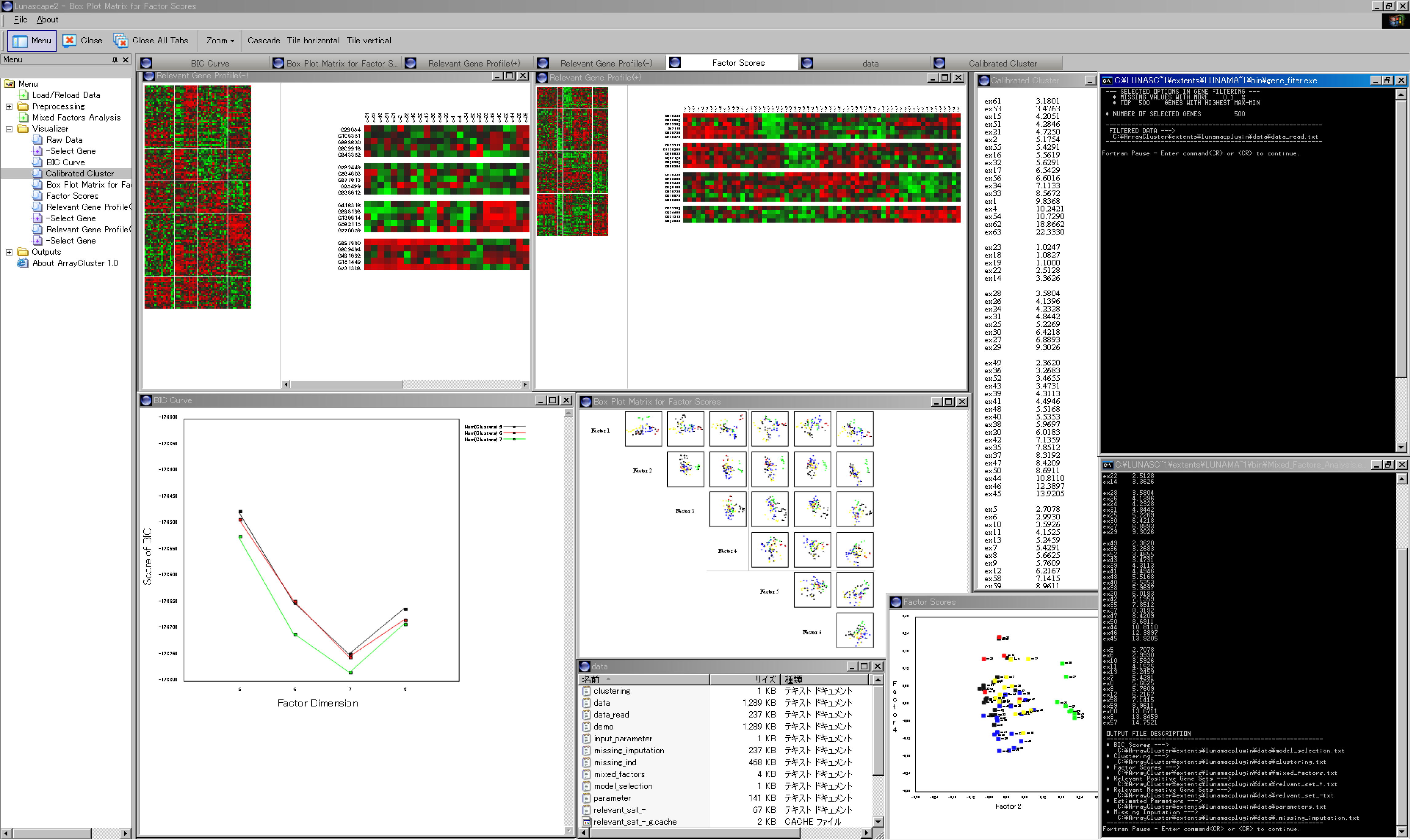
This work was carried out in the laboratory of Tomoyuki Higuchi, The Institute of Statistical Mathematics, Research Organization of Information and Systems, and the laboratory of
Satoru Miyano, DNA Information Analysis, Human Genome Center, Institute
of Medical Science, University of Tokyo.
Developers; Tomoyuki Higuchi, Ryo Yoshida, Seiya Imoto, Satoru Miyano
Copyright; (C) 2005- Tomoyuki Higuchi (C) 2005- Ryo Yoshida (C) 2005- Seiya Imoto (C) 2005-
Satoru Miyano (C) 2005- The Institute of Statistical Mathematics, Research Organization of Information and Systems (C) 2005- Human Genome
Center, Institute of Medical Science, University of Tokyo
